First accredited laboratory worldwide to offer nanopore sequencing of bacteria
The Institute for Infectious Diseases (IFIK) of the University of Bern is the first accredited laboratory worldwide to offer nanopore sequencing for the identification of bacteria.
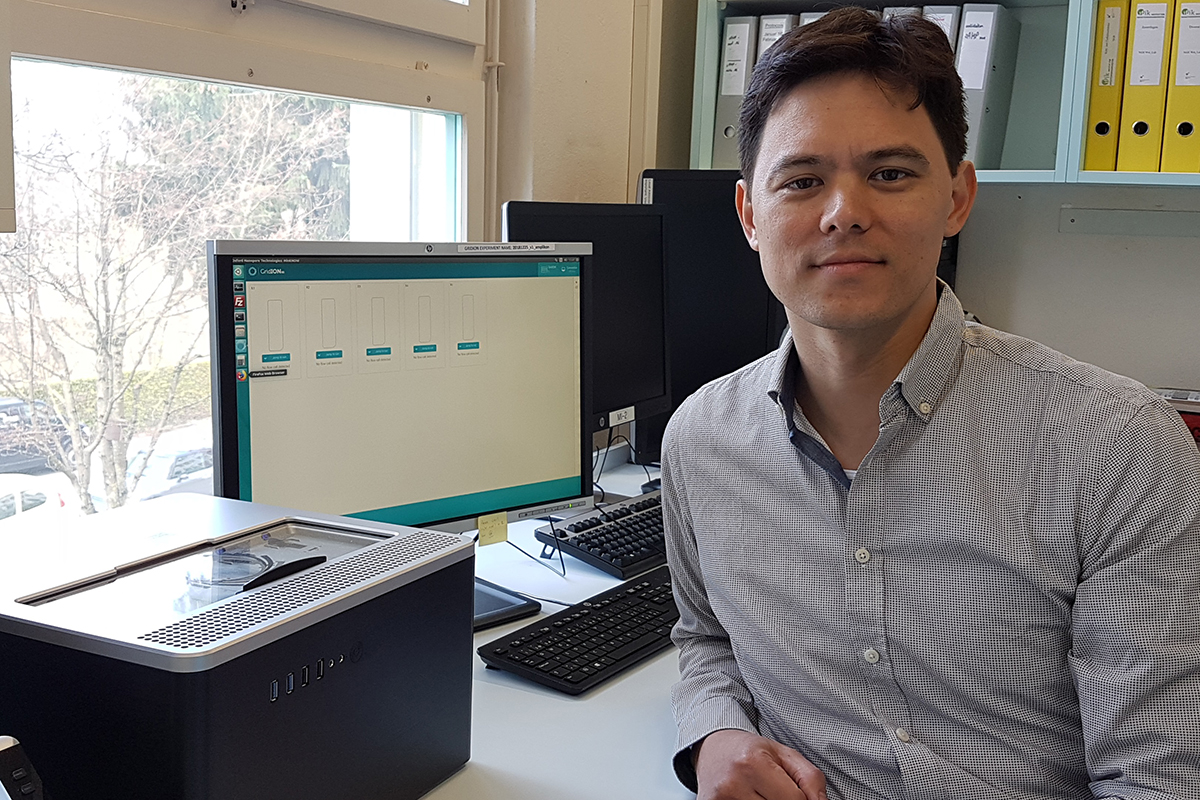
As of January 21, 2019, the Institute for Infection Diseases (IFIK) of the University of Bern is proud to be the first laboratory worldwide to have received the ISO/IEC 17025:2018 accreditation status to sequence specific DNA fragments from bacterial cultures using Oxford Nanopore Technologies devices in a medical diagnostic setting. The new sequencing technology enables to taxonomically identify pathogens more rapidly and cost-effectively. The IFIK Sequencing laboratory also obtained in mid-October 2018 the qualification of certified Nanopore Service Provider, the first one in Switzerland. Dr. Alban Ramette, Head Bioinformatics/Biostatistics group at the Institute for Infectious Diseases, tells «uniaktuell» how the technology works and how he will use it.

What does the accreditation mean for the Institute for Infectious Diseases (IFIK)?
Alban Ramette: To be the first laboratory worldwide to have received the accreditation to sequence specific DNA fragments using Oxford Nanopore Technologies devices is a great honor for us. This recognition puts Bern on the map as a center of excellence for health science, and IFIK as a place where game-changing technological innovations are efficiently translated to medical diagnostic applications.
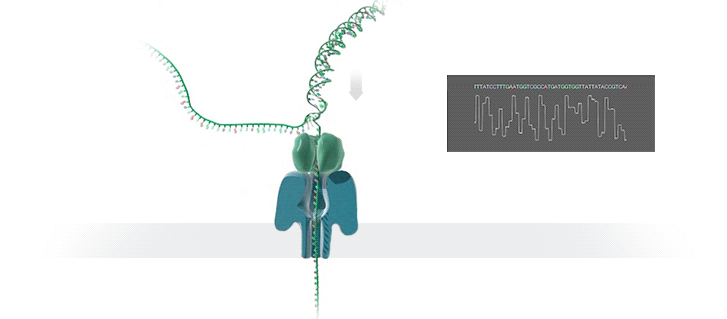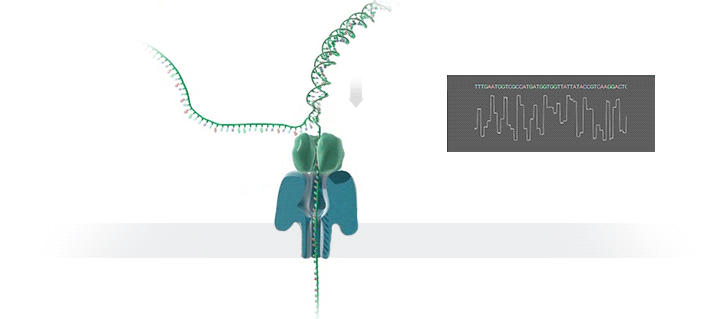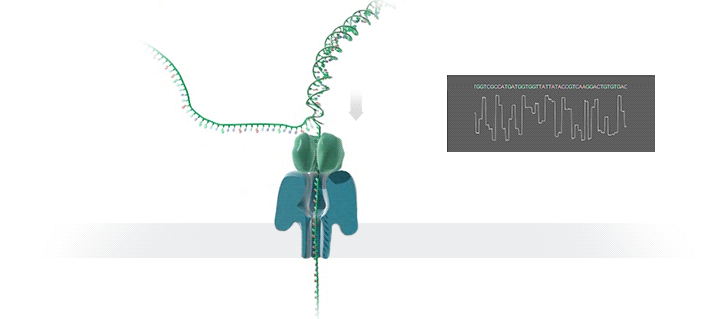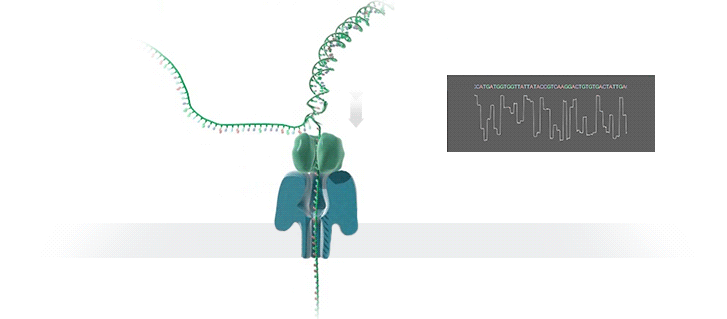
What advantage does the new technology offer, and how does it work?
The new technology enables fast sequencing of DNA fragments, and is ideal for medium-throughput assays where flexibility in terms of number of samples and short turn-around time is needed. Inside the so-called GridION device, an ionic current passes through nanopores (nano-scale holes puncturing membranes). Changes in current as biological molecules pass through the nanopore are recorded and used to identify the composition of the molecules. Because molecules are sequenced individually, the presence of molecules of different origins (for instance from a mixture of pathogenic and commensal bacteria) is readily identified and further addressed by matching individual sequences to their most likely reference sequences in bioinformatic databases.
What exactly are you analyzing?
When an uncommon or yet undescribed pathogen is isolated from a patient sample, we sequence common housekeeping genetic markers to identify the pathogen. The comparison of those signature sequences with those of known reference sequences enables a precise identification of the micro-organism at hand. This information is combined with the results of other diagnostic tests to obtain the most accurate identification of pathogenic bacteria, which is then communicated to clinicians.
Who might benefit from it?
Ultimately, patients will benefit from this technological improvement, as rapid and thorough analyses of samples sent to IFIK for clinical diagnostic purposes are key in rapidly providing the most accurate information available to treating physicians.
ABOUT THE PERSON

Dr. Alban Ramette, PhD Nat. Sc., is Head of Bioinformatics/Biostatistics group and of the sequencing platform at the Institute for Infectious Diseases (IFIK), University of Bern.
Contact:
Dr. Alban Ramette, PhD
Head Bioinformatics/Biostatistics group
Institute for Infectious Diseases, University of Bern, Switzerland
Tel.: +41 31 632 95 40 / alban.ramette@ifik.unibe.ch
The Institute for Infectious Diseases (IFIK)
The Institute for Infectious Diseases is the only university institute in Switzerland to combine all microbiological specialties (such as parasitology, bacteriology or virology) under one roof in teaching, research and diagnostics. Its research focuses on the development of antibiotic resistance, infections of the central nervous system as well as health and infections of intestinal bacteria.
ABOUT THE AUTHOR
Nathalie Matter is an editor and the "health & medicine" representative at the University of Bern Communication & Marketing Office.